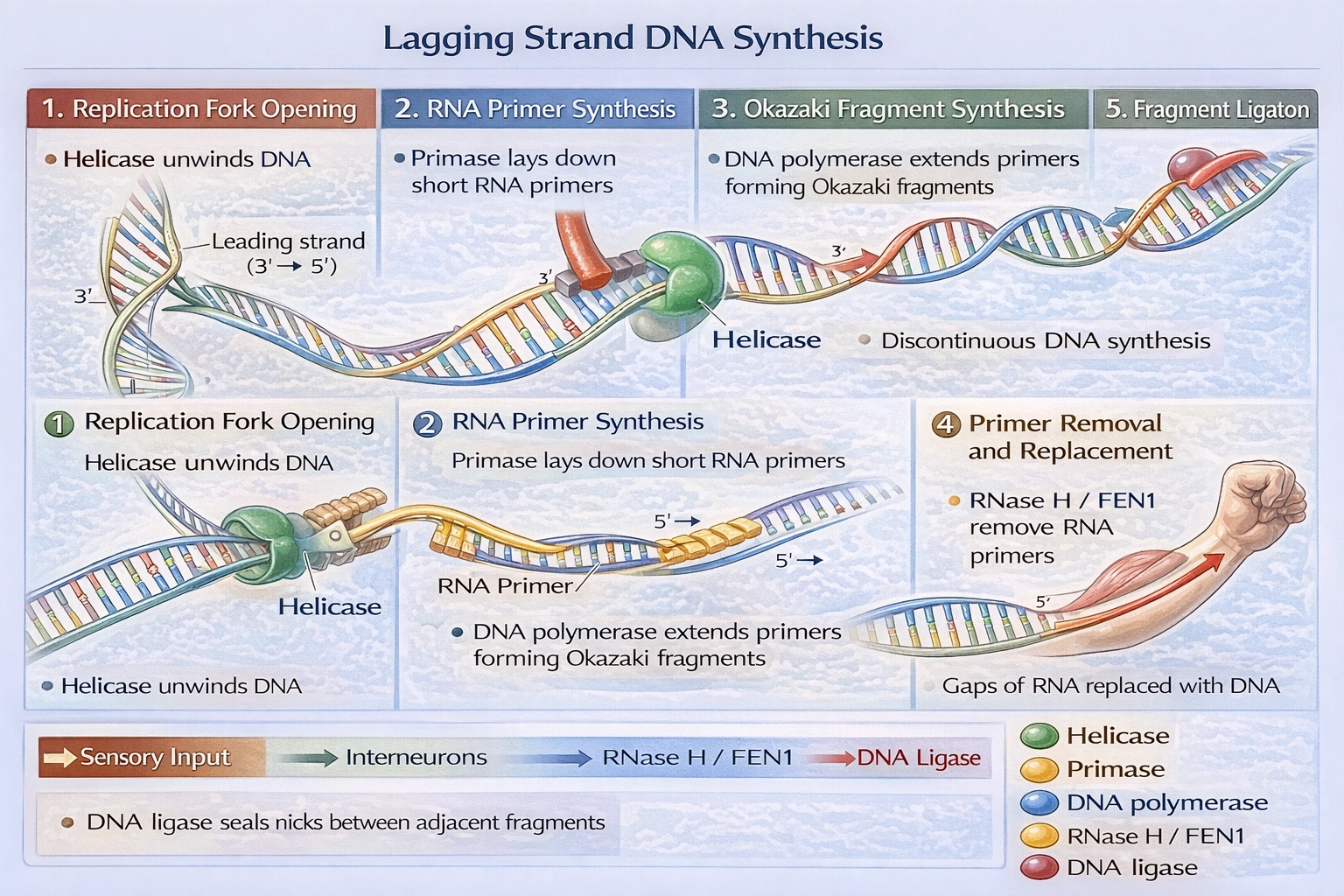
Describe the synthesis of the lagging strand of DNA.
-
A
DNA polymerases synthesize DNA continuously after initially attaching to a primase.
-
B
DNA polymerases synthesize DNA discontinuously in pieces called Okazaki fragments after initially attaching to primases.
-
C
DNA polymerases synthesize DNA discontinuously in pieces called Okazaki fragments after initially attaching to RNA primers.
-
D
DNA polymerases synthesize DNA discontinuously in pieces called Okazaki fragments which are joined together in the end by a DNA helicase.
DNA polymerases synthesize DNA discontinuously in pieces called Okazaki fragments after initially attaching to RNA primers.
During DNA replication, the lagging strand undergoes discontinuous synthesis because DNA polymerases can only add nucleotides in the 5'→3' direction while the replication fork opens in the opposite orientation relative to this strand. The enzyme primase (a specialized RNA polymerase) first synthesizes short RNA primers approximately 10–12 nucleotides long at intervals along the lagging strand template. DNA polymerase III in prokaryotes (or DNA polymerase δ in eukaryotes) then binds to the 3'-hydroxyl terminus of each RNA primer—not to primase itself—and extends the primer by adding deoxyribonucleotides in the 5'→3' direction away from the replication fork, producing Okazaki fragments typically 1,000–2,000 nucleotides long in prokaryotes and 100–200 nucleotides long in eukaryotes. After synthesis of adjacent fragments, RNase H and FEN1 (Flap Endonuclease 1) remove the RNA primers, DNA polymerase I (in prokaryotes) or DNA polymerase δ (in eukaryotes) fills the resulting gaps with DNA nucleotides, and DNA ligase catalyzes phosphodiester bond formation to covalently join the fragments into a continuous strand.
A) DNA polymerases synthesize DNA continuously after initially attaching to a primase
This description characterizes leading strand synthesis, not lagging strand synthesis. The leading strand is synthesized continuously in the 5'→3' direction toward the replication fork after a single RNA primer is laid down near the origin of replication. Furthermore, DNA polymerases do not "attach to primase"; primase is an enzyme that synthesizes RNA primers and then dissociates. DNA polymerases bind to the 3'-OH end of the RNA primer itself to initiate DNA synthesis. The terminology "attaching to a primase" misrepresents the molecular interaction and incorrectly applies continuous synthesis to the lagging strand.
B) DNA polymerases synthesize DNA discontinuously in pieces called Okazaki fragments after initially attaching to primases
While this option correctly identifies discontinuous synthesis and Okazaki fragments, it contains a critical terminology error: DNA polymerases attach to RNA primers, not to "primases." Primase (singular) is the enzyme complex responsible for synthesizing RNA primers; it is not a structural attachment site for DNA polymerases. After primase synthesizes a short RNA segment and dissociates from the template, DNA polymerase recognizes and binds to the free 3'-hydroxyl group of the RNA primer to begin elongation. Referring to "primases" as attachment points confuses the enzymatic catalyst (primase) with its product (RNA primer), representing a fundamental misunderstanding of replication biochemistry.
C) DNA polymerases synthesize DNA discontinuously in pieces called Okazaki fragments after initially attaching to RNA primers
This option accurately describes the molecular mechanism of lagging strand synthesis. Primase synthesizes discrete RNA primers at regular intervals along the exposed lagging strand template as the replication fork progresses. Each RNA primer provides a free 3'-OH group required by DNA polymerases to initiate DNA synthesis, as these enzymes cannot start synthesis de novo. DNA polymerase then extends each primer to form an Okazaki fragment. The discontinuous nature arises because each fragment must be initiated separately as new template becomes available with fork progression. This process requires coordinated action of multiple enzymes: primase for primer synthesis, DNA polymerase for elongation, RNase H/FEN1 for primer removal, and DNA ligase for fragment joining—collectively ensuring high-fidelity duplication of the entire genome despite the antiparallel nature of DNA and unidirectional activity of polymerases.
D) DNA polymerases synthesize DNA discontinuously in pieces called Okazaki fragments which are joined together in the end by a DNA helicase
This option correctly identifies discontinuous synthesis and Okazaki fragments but erroneously attributes fragment ligation to DNA helicase. DNA helicase (such as DnaB in prokaryotes or MCM complex in eukaryotes) functions exclusively to unwind the double-stranded DNA helix ahead of the replication fork by breaking hydrogen bonds between base pairs, using ATP hydrolysis to translocate along the DNA. Helicase plays no role in joining DNA fragments. The enzyme responsible for covalently sealing nicks between adjacent Okazaki fragments is DNA ligase, which catalyzes the formation of phosphodiester bonds between the 3'-OH end of one fragment and the 5'-phosphate end of the next fragment after RNA primer replacement. Confusing helicase with ligase misattributes fundamentally distinct enzymatic functions within the replication machinery.
Conclusion:
Lagging strand synthesis exemplifies an elegant biochemical solution to the topological constraints imposed by DNA's antiparallel structure and the unidirectional 5'→3' activity of DNA polymerases. The process requires repeated cycles of RNA priming by primase, discontinuous DNA elongation producing Okazaki fragments, precise RNA primer excision, gap filling with DNA nucleotides, and final ligation by DNA ligase—not helicase—to generate a continuous daughter strand. Option C correctly identifies RNA primers as the essential initiation points for DNA polymerase activity and accurately describes the discontinuous nature of lagging strand replication, while the other options contain critical errors regarding synthesis continuity, molecular attachment sites, or enzymatic functions responsible for fragment joining.